This is an old revision of the document!
Table of Contents
MCULE COMPOUND SOURCING USER GUIDE
This guide summarizes the major steps of the compound selection and ordering process in Mcule. If you have any questions, feel free to contact us at support@mcule.com.
Basic compound selection
The Purchasable compounds collection of Mcule contains over 6 million unique structures and 10 million purchasable products. To keep our database as up-to-date as possible we have developed batch and live handlers that can automatically update stock availability information for millions of products per day. This makes the Mcule database the most up-to-date database of screening compounds currently available. Our compound registration system (Mcule Advanced Curation) includes over 80 steps and 150 substeps including normalization, preparation steps and structural checks. This unique registration system filters out the majority of problematic structures including drawing errors and stereochemical problems. To search this large chemical space we have developed efficient and fast searching tools to help you finding the right compounds for your drug discovery project. Below we list the various searching options you can select in Mcule.
Find Chemicals / Single query
You can use this option to quickly find specific compounds/chemicals or close analogs of a single query compound in the Mcule database.
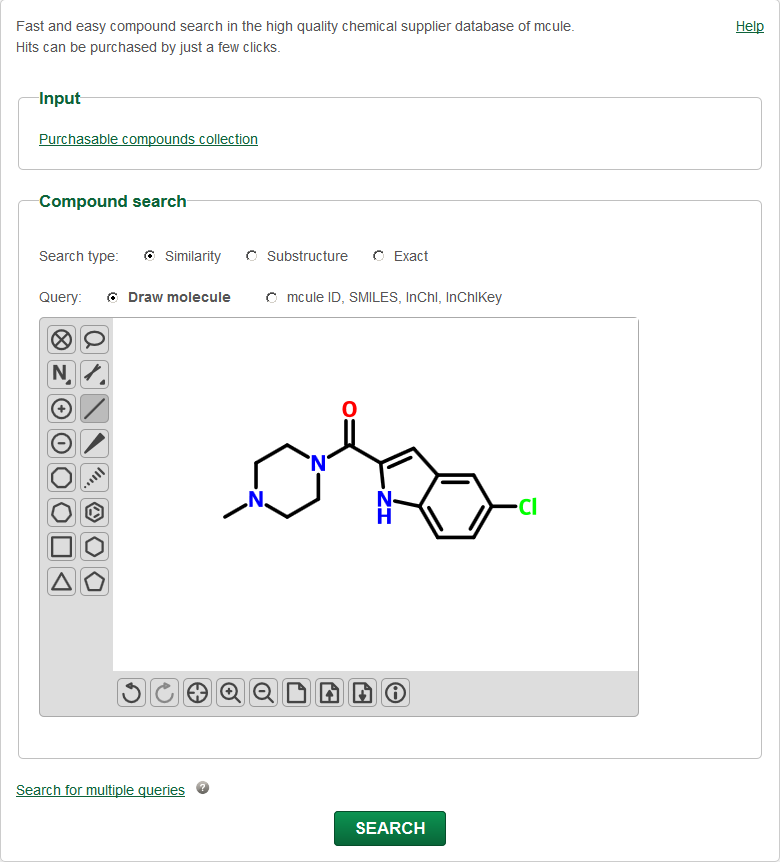 Exact, Similarity and Substructure searches.
Exact, Similarity and Substructure searches.
Search by SMILES, InChI, InChIKey, Mcule ID or use ChemWriter (Javascript drawer) to specify your query.
Searching in over 6M purchasable compounds (Mcule database).
Find Chemicals / Multiple queries exact
https://mcule.com/search/multi
With this option you can run exact searches with multiple query molecules. It can be useful for ordering a set of ligands e.g. virtual screening hits. You can paste or upload structures in multiple formats, such as Mcule IDs, InChI or SMILES strings, InChIKeys and even SDF files and use them as queries in the search.
Find Chemicals / Multiple queries similarity
https://mcule.com/search/screen/?template=multiple-queries-similarity
If you have multiple hits containing common structural elements (e.g. multiple analogs of the same scaffold), it makes sense to include all your hits in a single search as multiple queries. Depending on the scoring method used, the searching algorithm will retrieve hits that are most similar to your query molecules. Best scoring method will retrieve the similarity score to the most similar query molecule. Average scoring method will calculate an average similarity score to all query molecules.
Find Chemicals / Mcule ID list
https://mcule.com/collection/create/from-ids
Searching for Mcule IDs is the most straightforward and effective way to retrieve a set of molecules from the Mcule database. For example, if you have downloaded the Purchasable compounds collection from our Database Download page, and you want to come back for purchasing some compounds, you can paste Mcule IDs in the below form. All exported SDF files that you download from Mcule contain the corresponding Mcule IDs as a separate SDF field. If you have both a list of Mcule IDs and SDF, it is preferable to use the Mcule ID to retrieve compounds, as it more clearly identifies the structures.
Smart compound selection
https://mcule.com/search/screen
The Mcule technology revolutionizes compound selection either for virtual screening or for screening library design. The Workflow Builder can be used to build custom screening and filtering workflows by putting together powerful and advanced cheminformatic and molecular modeling tools like LEGO bricks. It can be used to run structure-based and ligand-based virtual screening workflows and the best hits can be purchased by just a few clicks. It enables the design of screening libraries based on various properties including chemical diversity (large-scale diversity selection) and supplier information (e.g. purity, available stock amount, delivery time, etc.).
Compound collection management
In Mcule you can select particular compounds either manually or by using the basic and advanced (smart) searching tools and store them in collections. The owner of a collection can set its privacy level to Private (only users with permission can get access), Unlisted (only people who know the specific URL of the collection can get access), or Public (accessible for everybody). Your search results (set of molecules and associated information) are also automatically stored in collections. Not only the search results but the queries are saved as well, so that you can go back and refine your queries until you get the optimal search results. Molecule collection can be stored, deleted, modified, merged, imported, exported and shared with your colleagues. Collections can be visualized in multiple ways together with associated data such as various properties, search and calculation results. Tables can be quickly sorted by these properties. Uploaded compounds are also stored in collections and can be used as input for any searches (all associated data incl. phys-chem properties will be automatically generated).