This is an old revision of the document!
Search
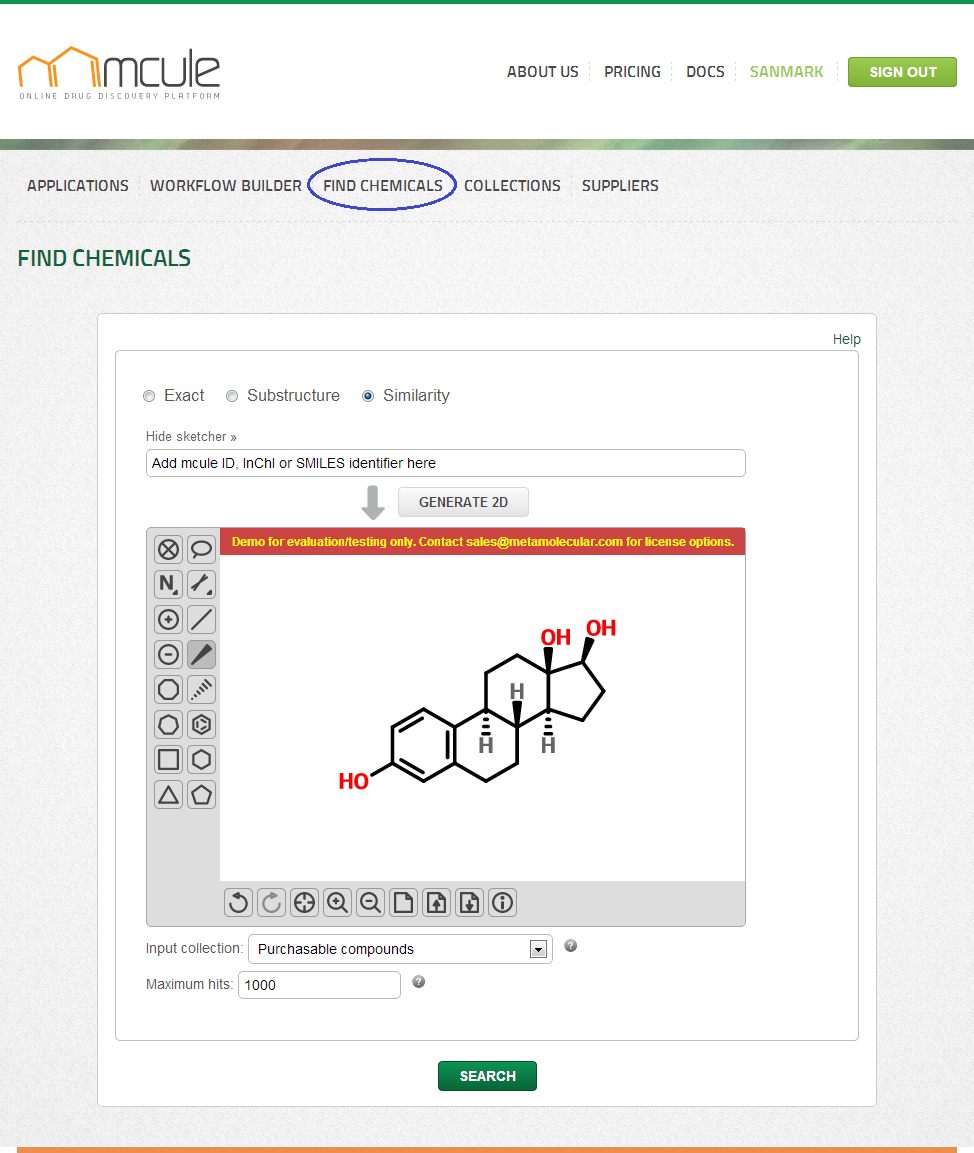
Under the “Search” tab three simple search types are available: “Exact”, “Similarity” and “Substructure” searches.
“Exact search” can be useful when looking for a specific molecule (e.g. a reference ligand).
“Similarity search” can retrieve close analogs of a query molecule (e.g. an HTS hit). “Similarity threshold” (Tanimoto coefficient) controls how similar the hits from the query need to be.
“Substructure search” can be used to find all molecules that contain a specific substructure (e.g. the main scaffold of a reference ligand).
Queries can be drawn by the molecule sketcher, which can be hidden and reopened by clicking on the “Hide/Show sketcher” link. Queries can be also defined by entering mcule IDs, InChI or SMILES strings into the input field. 2D representations can be generated by clicking on the “Generate 2D” button. Note: If the “Generate 2D” button was clicked, the search will be performed on the generated 2D representation (2D SDF exported from the sketcher). If, however, the “Generate 2D” button was not clicked, the search will be directly performed on the mcule ID, InChI or SMILES strings coming from the input field. The results might be slightly different in these two cases.
You can select on which collection the search will be performed in the “Input collection” field. By default, the “Purchasable compounds collection” is set as input collection. Other collections (e.g. public collections or your previously created collections) can be also selected.
The “Maximum hits” field is set to 1,000 by default. If you don't want to limit the number of results, you can delete the number from this field. Note that if you are using the Free package, the maximum number of molecules in a single collection is 10,000, which implicitly limits the “Maximum hits” to 10,000.